1. Introduction
For much of modern toxicology, the evaluation of chemical safety has relied—almost unquestioningly—on in vivo experimentation. Animal models, particularly rodents, have served as the foundational systems through which hazard identification and dose–response relationships are inferred. And yet, as indispensable as these models once seemed, their limitations have become increasingly difficult to ignore. The sheer scale of contemporary chemical production—where thousands of new compounds are introduced annually—has exposed a structural mismatch between testing capacity and regulatory demand (National Research Council, 2007). Traditional in vivo studies, while methodologically rigorous, are slow, costly, and ethically burdensome. It is not uncommon for a single regulatory assessment to require years of experimentation, thousands of animals, and substantial financial investment, often without guaranteeing direct human relevance.
More troubling, perhaps, is the persistent issue of translational uncertainty. Biological responses observed in animal systems do not always align with human physiology, largely due to interspecies differences in metabolism, signaling pathways, and organ-specific sensitivities (Gottmann et al., 2001). This disconnect introduces a degree of ambiguity into risk assessment—one that becomes particularly consequential in drug development and environmental health decision-making. In this light, the long-standing “gold standard” begins to appear less like a definitive benchmark and more like a constrained approximation of biological reality.
It is within this context that toxicology has begun—gradually but decisively—to redefine itself. Over the past two decades, a conceptual shift has emerged, moving away from descriptive, endpoint-driven animal testing toward mechanism-based, human-relevant approaches. Central to this transformation is the integration of in vitro and in silico methodologies, which together promise not only higher throughput but also deeper mechanistic insight (Kavlock et al., 2009; Sun et al., 2012). High-throughput screening technologies, particularly quantitative high-throughput screening (qHTS), have enabled the rapid evaluation of thousands of compounds across diverse biological targets, generating datasets of unprecedented scale and resolution (Shukla et al., 2010).
At the same time, advances in computational toxicology have opened new avenues for predictive modeling. Early efforts in this domain—such as quantitative structure–activity relationship (QSAR) modeling—were grounded in the premise that chemical structure encodes biological activity (McKinney, 1985). While promising, these models were often constrained by limited data quality and simplistic assumptions (Cronin & Schultz, 2003). However, with the advent of machine learning and improved data curation practices, contemporary models are increasingly capable of capturing nonlinear relationships and multidimensional interactions within toxicological systems (Zhu et al., 2008; Merlot, 2010).
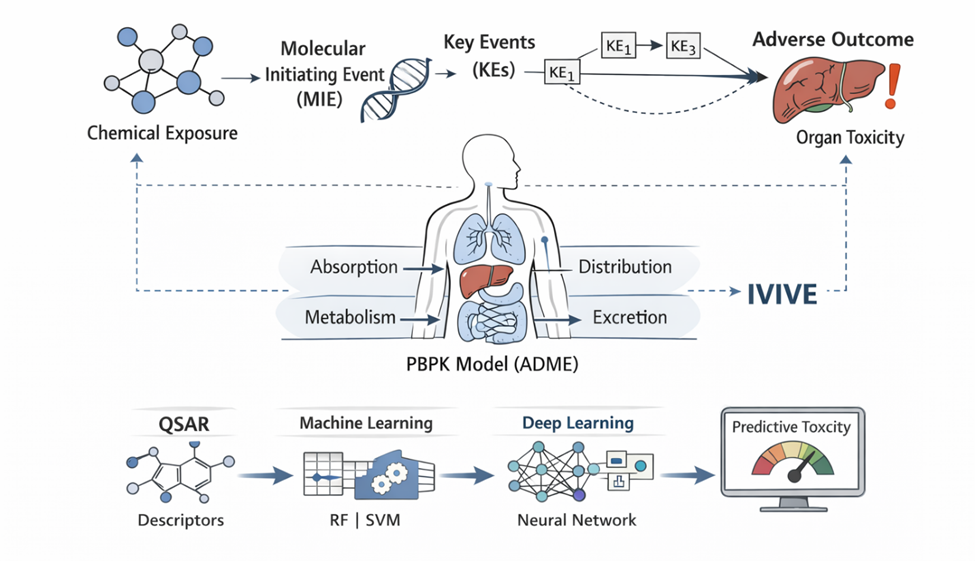
What is emerging, then, is not merely a replacement of one methodology with another, but rather a convergence—a synthesis of diverse data streams into what might be termed “foundation toxicity models.” These models aim to integrate chemical descriptors, in vitro assay outputs, and in vivo outcomes into unified predictive frameworks. In doing so, they attempt to bridge a longstanding gap: the disconnect between molecular-level perturbations and organism-level adverse effects. This challenge—often referred to as the in vitro to in vivo extrapolation (IVIVE) problem—remains one of the central obstacles in modern toxicology (Thomas et al., 2013).
The development of mechanistic frameworks such as Adverse Outcome Pathways (AOPs) has provided a conceptual scaffold for addressing this gap. AOPs map the progression from a molecular initiating event through a series of key biological events to an adverse outcome at the organism or population level (Ankley et al., 2010). In principle, this structured representation allows for the alignment of in vitro assay data with in vivo toxicity endpoints, thereby enhancing both interpretability and predictive accuracy. Yet, despite their conceptual elegance, AOP-based models still face challenges in terms of completeness, validation, and integration with quantitative modeling approaches.
It is worth noting that this shift toward integrative modeling is not occurring in isolation. Large-scale collaborative initiatives—such as the Tox21 program—have played a pivotal role in accelerating methodological innovation and data sharing (Schmidt, 2009). These efforts reflect a broader recognition that the complexity of toxicological systems cannot be adequately addressed through isolated experimental approaches. Instead, what is required is a systems-level perspective—one that embraces heterogeneity, leverages computational power, and remains grounded in biological plausibility.
Historically, the roots of this transformation can be traced back several decades. Early computational efforts, including expert systems and predictive toxicology evaluation projects, hinted at the possibility of in silico hazard prediction (Benfenati & Gini, 1997; Bristol et al., 1996). Similarly, foundational in vitro assays—such as neutral red uptake and colorimetric viability tests—demonstrated that cellular responses could serve as proxies for organismal toxicity (Borenfreund & Puerner, 1985; Mosmann, 1983). While these methods were initially limited in scope, they laid the groundwork for the high-throughput, data-rich paradigms that define contemporary toxicology.
Still, the path forward is not without uncertainty. The integration of heterogeneous data—spanning chemical space, biological systems, and temporal scales—introduces both technical and conceptual challenges. Data quality remains a persistent concern, as inconsistencies in experimental design and reporting can undermine model reliability (Gottmann et al., 2001). Moreover, the selection of appropriate molecular descriptors and statistical algorithms requires careful consideration, particularly given the risk of overfitting and reduced generalizability (Cronin & Schultz, 2003).
And yet, despite these challenges, the trajectory is unmistakable. Toxicology is moving toward a future in which predictive models—grounded in mechanistic understanding and supported by diverse data streams—play a central role in risk assessment. Foundation toxicity models, in this sense, represent both a culmination of past efforts and a starting point for new methodological innovations.
This review, therefore, seeks to examine the evolution and current state of these integrative approaches. Specifically, it explores how in vivo, in vitro, and chemical knowledge can be harmonized within unified predictive frameworks; evaluates the role of machine learning and molecular descriptors in enhancing model performance; and considers the extent to which mechanistic frameworks such as AOPs can improve IVIVE. In doing so, it aims not only to synthesize existing knowledge but also to reflect—perhaps cautiously—on the future direction of toxicity testing in an increasingly data-driven scientific landscape.