1. Introduction
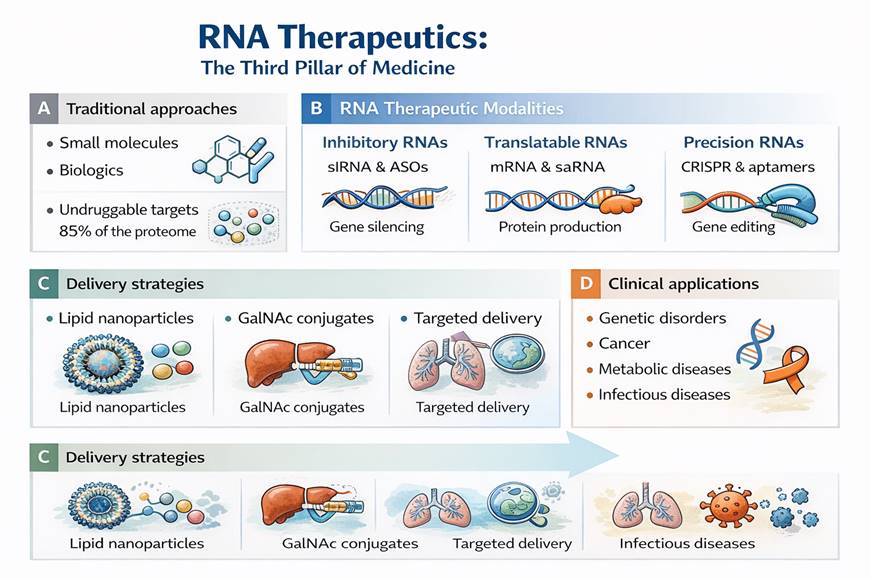
Over the past decade, RNA-based therapeutics have moved from theoretical promise to clinical reality, redefining the landscape of modern drug development. Increasingly described as the “third pillar” of therapeutics—alongside small molecules and biologics—RNA medicines represent a conceptual shift in how disease is approached and treated (Al-Hamdani et al., 2023; Shahid, 2025). Traditional pharmacology largely depends on modulating protein function through binding to accessible active sites. Yet, it is estimated that nearly 85% of the human proteome lacks such druggable pockets, leaving many genetic drivers of disease beyond the reach of conventional approaches (Jones et al., 2024; Paunovska et al., 2022). RNA-based platforms overcome this limitation by acting upstream of protein synthesis, enabling direct modulation of gene expression at the transcript level. In principle, once a pathogenic gene sequence is identified, it becomes a potential therapeutic target.
This systematic review synthesizes current advances across the diverse modalities of RNA therapeutics, with particular emphasis on their mechanisms of action, clinical maturity, and delivery technologies that underpin translational success. By organizing evidence across inhibitory, translatable, and precision RNA platforms, we aim to provide a coherent framework for understanding how RNA medicines are reshaping treatment paradigms across genetic, metabolic, oncologic, and infectious diseases.
Among RNA therapeutics, inhibitory modalities are the most clinically established. Antisense oligonucleotides (ASOs) are short, synthetic, single-stranded nucleic acid sequences designed to bind complementary RNA transcripts through Watson–Crick base pairing (Shahid, 2025). Their mechanisms are diverse. Some ASOs recruit RNase H1, leading to selective degradation of the target mRNA, while others function sterically—modulating pre-mRNA splicing or blocking translation without transcript destruction (Fontanellas et al., 2025; Torrisi et al., 2026). The approval of nusinersen for spinal muscular atrophy marked a pivotal milestone for this field, demonstrating that rational splice modulation can restore functional protein production and alter the natural history of a devastating genetic disease (Shahid, 2025; Torrisi et al., 2026).
Similarly, small interfering RNAs (siRNAs) harness the endogenous RNA interference (RNAi) pathway. After cellular entry, siRNAs are incorporated into the RNA-induced silencing complex (RISC), which uses sequence complementarity to guide catalytic cleavage of target mRNA (Musa et al., 2024; Shevelev et al., 2025). Clinically approved siRNA therapeutics such as patisiran and inclisiran illustrate the potency and durability of this mechanism. Chemical stabilization strategies—such as backbone modifications and targeted conjugation—enable prolonged gene silencing with infrequent dosing schedules.
MicroRNA (miRNA) modulators extend this paradigm further. Unlike siRNAs, miRNAs exhibit partial complementarity and may regulate entire gene networks rather than single transcripts. This broader regulatory scope offers therapeutic opportunities in complex diseases but also introduces challenges related to specificity and off-target effects (Banskota et al., 2022; Volpini et al., 2023). Collectively, inhibitory RNA modalities demonstrate how precise transcript-level intervention can translate into durable clinical benefit.
In contrast to inhibitory platforms, translatable RNAs function by delivering genetic instructions that enable cells to produce therapeutic proteins. Messenger RNA (mRNA) therapeutics are perhaps the most visible example of this strategy. By introducing synthetic mRNA directly into the cytoplasm, cells are temporarily reprogrammed to synthesize proteins, antigens, or enzymes without the risks associated with genomic integration (Khan et al., 2025; Shahid, 2025). The transient nature of mRNA expression enhances safety, while advances in nucleoside modification and structural optimization improve translational efficiency and reduce innate immune activation (Song et al., 2025).
To address limitations in expression duration, researchers have developed self-amplifying mRNA (saRNA). By incorporating viral replicase elements, saRNA constructs replicate intracellularly, generating multiple RNA copies from a single administered dose (Ho et al., 2025; Wang et al., 2025). This amplification allows sustained protein production at substantially lower doses, potentially improving cost-effectiveness and reducing adverse effects.
Circular RNA (circRNA) represents another emerging platform. Unlike linear mRNA, circRNA molecules are covalently closed loops lacking free 5' and 3' ends, rendering them resistant to exonuclease-mediated degradation (Dalabehera et al., 2025; Shahid, 2025). Engineering strategies—such as incorporation of internal ribosome entry sites (IRES)—enable efficient cap-independent translation. The enhanced stability of circRNA suggests potential advantages for chronic conditions requiring prolonged therapeutic protein expression (Fontanellas et al., 2025). Together, these encoding RNA modalities expand the therapeutic toolbox from gene silencing to controlled protein synthesis.
Beyond silencing and protein production, RNA also serves as a guide and targeting scaffold in precision medicine. CRISPR–Cas systems rely on guide RNAs (gRNAs) to direct nucleases to specific genomic loci, enabling permanent gene disruption or correction (Chakraborty et al., 2026). The clinical approval of CRISPR-based therapies for genetic disorders highlights the transformative potential of RNA-guided genome editing. Unlike transient RNA interference, CRISPR-mediated interventions may offer durable, possibly curative outcomes.
RNA aptamers add another dimension to precision therapeutics. These single-stranded oligonucleotides fold into defined three-dimensional conformations capable of binding proteins or cell-surface receptors with high specificity (Zhu et al., 2022). Often described as “chemical antibodies,” aptamers can function as direct antagonists or as targeting ligands to guide other therapeutic payloads. Their synthetic nature, lower immunogenicity, and tunable pharmacokinetics make them attractive complements to antibody-based therapies.
Despite conceptual elegance, RNA therapeutics face formidable biological barriers. Naked RNA molecules are inherently unstable, rapidly degraded by ubiquitous ribonucleases (RNases), and prone to immune detection through Toll-like receptors (TLRs) (Khan et al., 2025; Shahid, 2025). Even when internalized via endocytosis, only a small fraction—estimated at 1% to 2%—escapes the endosome to reach the cytoplasm, where therapeutic action occurs (Navid Talemi et al., 2026; Shahid, 2025). This “endosomal escape” bottleneck remains a central obstacle in the field.
Lipid nanoparticles (LNPs) have emerged as the current gold standard for systemic RNA delivery. These multi-component systems typically include ionizable lipids, cholesterol, helper phospholipids, and polyethylene glycol (PEG)-lipids (Navid Talemi et al., 2026; Paunovska et al., 2022). Ionizable lipids are engineered to remain neutral at physiological pH but acquire positive charge in the acidic endosomal environment, promoting membrane destabilization and cytoplasmic release of RNA cargo (Navid Talemi et al., 2026; Shahid, 2025). While highly effective for hepatic targeting and vaccine applications, LNPs exhibit intrinsic liver tropism, limiting efficient delivery to extrahepatic tissues (Dalabehera et al., 2025; Khan et al., 2025).
Recognizing these limitations, current research is focused on expanding tissue specificity and improving precision targeting. Selective Organ Targeting (SORT) technology modifies conventional LNP formulations by incorporating “tuning” lipids that alter biodistribution patterns (Dalabehera et al., 2025; Navid Talemi et al., 2026). Through rational compositional changes, RNA payloads can be preferentially directed to the lungs, spleen, or other organs, advancing the concept of “mRNA Delivery 2.0.”
GalNAc conjugation remains the benchmark for hepatocyte-specific delivery, particularly for siRNA therapeutics administered subcutaneously (Fontanellas et al., 2025; Shahid, 2025). By exploiting the asialoglycoprotein receptor highly expressed on hepatocytes, GalNAc-conjugated RNAs achieve potent gene silencing with remarkable precision.
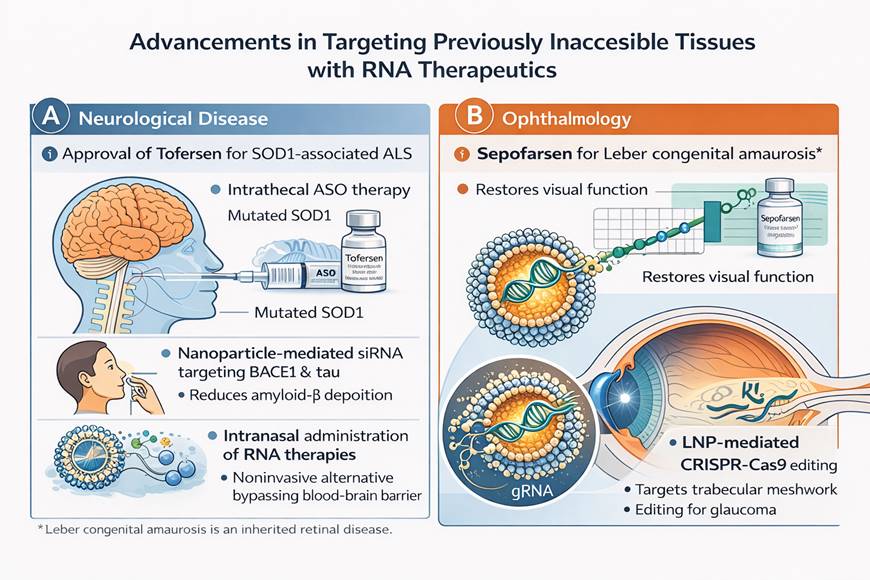
Alternative platforms, including extracellular vesicles (exosomes), polymeric nanoparticles (PNPs), and cell-penetrating peptides (CPPs), are under active investigation (Du et al., 2025; Gu et al., 2026; Setia et al., 2026). Exosomes, in particular, offer inherent biocompatibility and the capacity to traverse biological barriers such as the blood–brain barrier. Concurrently, artificial intelligence and machine learning are increasingly employed to predict biodistribution profiles and optimize lipid compositions, enabling data-driven design of next-generation delivery systems (Dalabehera et al., 2025; Navid Talemi et al., 2026).
Taken together, RNA therapeutics represent a versatile and rapidly evolving class of medicines capable of addressing previously intractable diseases. However, their clinical success depends not only on molecular design but also on overcoming complex delivery constraints. This systematic review integrates mechanistic, translational, and technological evidence to critically evaluate the current state of RNA-based therapeutics and identify key gaps that must be addressed to fully realize their transformative potential in precision medicine.